WebProtege
This tutorial gives an introduction to using WebProtégé for curating PGx recommendations.
Protégé is developed by Stanford University. And general-puropose tutorials can be found at the Stanford Protégé Wiki.
Contents
User interface of Pharmacoracle's WebProtégé
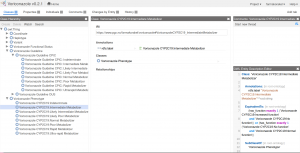
When opening an PGx guideline in WebProtégé for the first time, we are presented to the Classes tab, containing four so-called views (frames)
- Class Hierarchy (a tree of all class entities of this ontology)
- Class (for description and editing of each class entity)
- Comments (for chatting with other curators)
- Project Feed (to see changes to the project.)
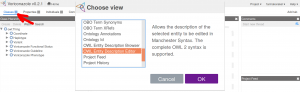
The user interface of WebProtégé can be modified. We suggest to replace the view Project Feed with the view OWL Entity Description Editor. In this way the editor/curator can inspect semantic properties of the class entities that are not shown in the default view. We suggest to remove the Project Feed in order to make the interface cleaner. The user can modify the interface as she or he pleases. Available views can be browsed by clicking on the three-bar symbol next to the Classes tab.
Using WebProtégé for curation of PGx recommendations
We suppose that the user has modified the Classes tab as explained above. We will use the drug voriconazole as an example. We have downloaded the CPIC guideline from the PharmGKB API and converted the PharmGKB JSON-LD file to the OWL format that can be edited in WebProtégé.
Curation of the functional status of star alleles
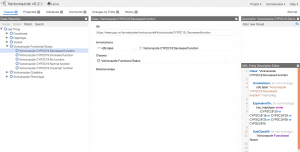
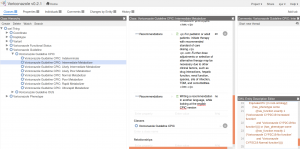
By navigating in the Class Hierachy view, the curator can e.g. view the class element Voriconazole CYP2C19:Decreased function. (The naming is based on the functional status that is provided by PharmGKB's API, and the drug and gene name is added because functional status is gene specific and possibly drug specific.) In the definition of the Voriconazole CYP2C19:Decreased function, pay attention to the OWL Entity Description Editor where we see that the Voriconazole CYP2C19:Decreased function is equivalent to a gene that has haplotype either CYP2C19*10, CYP2C19*16, CYP2C19*19, CYP2C19*25, CYP2C19*26 or CYP2C19*9.
Although the Functional Status can be manually modified by the curator, we believe that it is better to edit the Functional Status programmatically, based on the current PharmGKB functional status. By starting a discussion in the Comment view, suggestions can be included in the next version of the ontology.
Curation of the Phenotype of patient diplotypes
CPIC has recommended to use the term Phenotype for the metabolization status, transportation function or presence of a particular trait. Currently, this Phenotype is not a independent entity in the PharmGKB API. This means that for a drug that is affected by two genes, the Phenotype is aggregated for the two genes. We suggest to make a phenotype per gene, and make a recommendation for the combination of phenotypes. If more genes are added later, then it may be easier to expand the guideline. Another reason for modelling phenotypes on a per gene basis is that we want to make a guideline that is not drug-dependent, but rather made for families of drugs for Psychopharmaca or Cancer, and involving very many different genes.
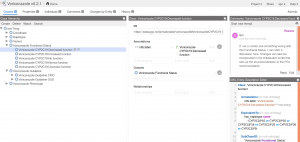
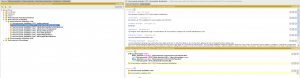
As an example, we see that the phenotype Voriconazole CYP2C19:Intermediate Metabolizer is either a heterozygote Voriconazole CYP2C19:Increased function/No function or a Voriconazole CYP2C19:No function/Normal function. As for the functional status, we suggest that curators start a discussion if definitions of Phenotypes should be changed.
Curation of the Recommendation
Finally, the WebProtégé can be used for writing the actual prose of the recommendations. For instance, if the curator wants to translate the CPIC recommendations into another language, in the Class view, she can enter a property Recommendations, a recommendation text and tagging it with the appropriate language.
Downloading the guideline in OWL format
One way to download a new revision of the guideline, is to go to the History tab, click on the revision number that you want to download, and download this as a .zip-file.
Try the Pharmacoracle WebProtégé yourself
Either contact us to register a new account at the WebProtégé, and for access to test ontologies
or try the Tutorial account:
user: Tutorial
password: Tutorial2018
The Tutorial account contains definitions of PGx alleles, PGx variants, functional status, phenotype and recommendations for the drug azathioprine.
Using Desktop Protégé for curation of PGx recommendations
Collaborative curation of PGx recommendations can quite easily be performed in WebProtégé, as explained above (of course, as with any tool, curators have to get used to it).
However, for quality assurance of PGx guideline semantics, we still need to use the desktop version of Protégé[1].
When changing the semantic structure of the guidelines, as explained for Phenotype in the section above, we need to be sure that our new guideline structure is equivalent to the structure we get from PharmGKB.
Consistency can be checked automatically in the Desktop Protege by running a reasoner (e.g. keyboard shortcut Ctrl-r). We then see if our structurally different OUS guidelines give the same recommendations as the CPIC guidelines.
- ↑ Support for semantic reasoning may be introduced in WebProtege at a later stage